Computes DPIT residuals for regression models with binary outcomes
using the observed responses (y) and their fitted distributional parameters(prob).
Usage
dpit_bin(y, prob, plot=TRUE, scale="normal", line_args=list(), ...)Arguments
- y
An observed outcome vector.
- prob
A vector of fitted probabilities of one.
- plot
A logical value indicating whether or not to return QQ-plot
- scale
You can choose the scale of the residuals among
normalanduniform. The sample quantiles of the residuals are plotted against the theoretical quantiles of a standard normal distribution under the normal scale, and against the theoretical quantiles of a uniform (0,1) distribution under the uniform scale. The default scale isnormal.- line_args
A named list of graphical parameters passed to
graphics::abline()to modify the reference (red) 45° line in the QQ plot. If left empty, a default red dashed line is drawn.- ...
Additional graphical arguments passed to
stats::qqplot()for customizing the QQ plot (e.g.,pch,col,cex,xlab,ylab).
Details
For formulation details on discrete outcomes, see dpit_pois.
Examples
## Binary example
n <- 500
set.seed(1234)
# Covariates
x1 <- rnorm(n, 1, 1)
x2 <- rbinom(n, 1, 0.7)
# Coefficients
beta0 <- -5
beta1 <- 2
beta2 <- 1
beta3 <- 3
q1 <- 1 / (1 + exp(beta0 + beta1 * x1 + beta2 * x2 + beta3 * x1 * x2))
y1 <- rbinom(n, size = 1, prob = 1 - q1)
# True model
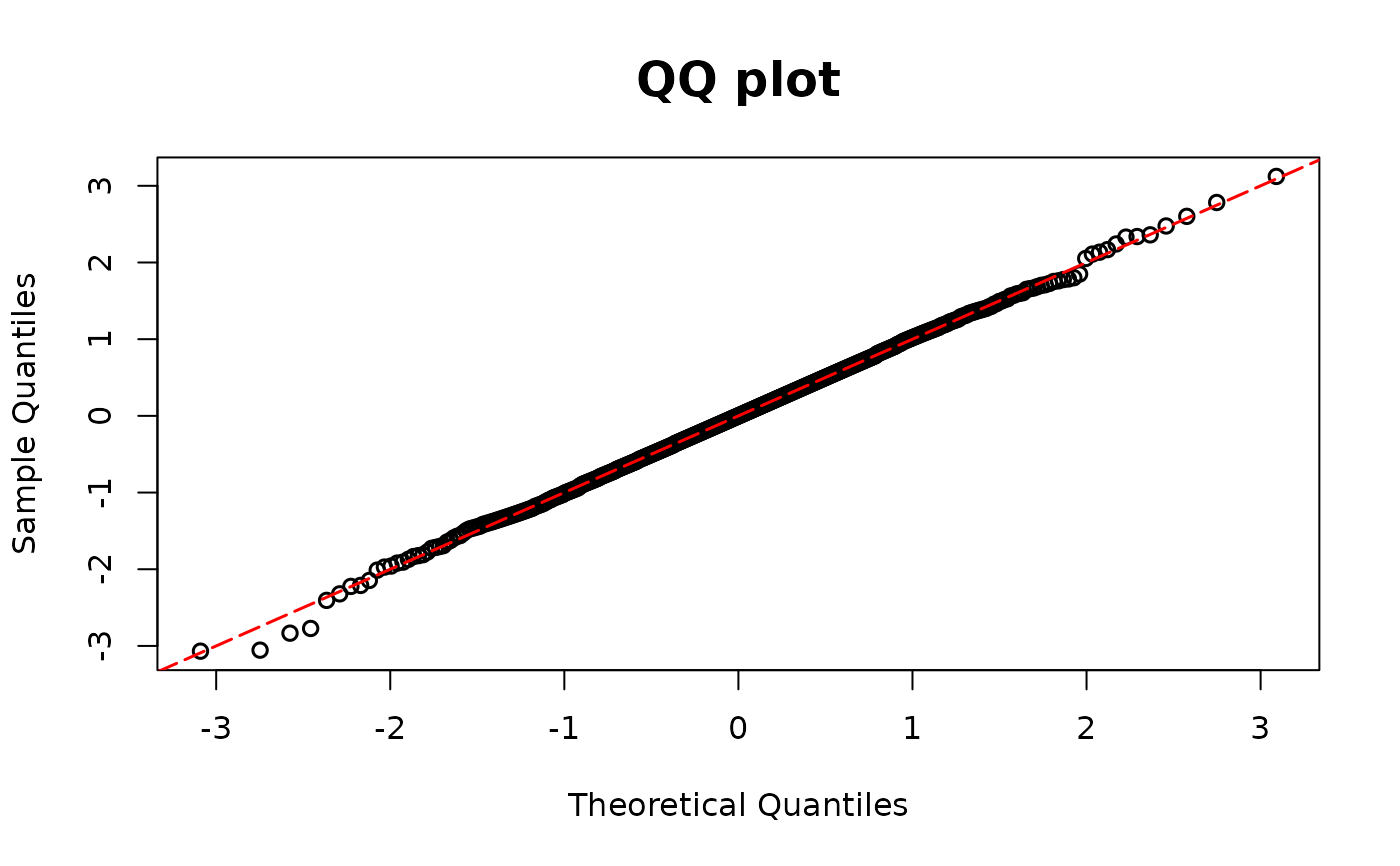
model01 <- glm(y1 ~ x1 * x2, family = binomial(link = "logit"))
fitted1 <- fitted(model01)
y1 <- model01$y
resid.bin1 <- dpit_bin(y=y1, prob=fitted1)
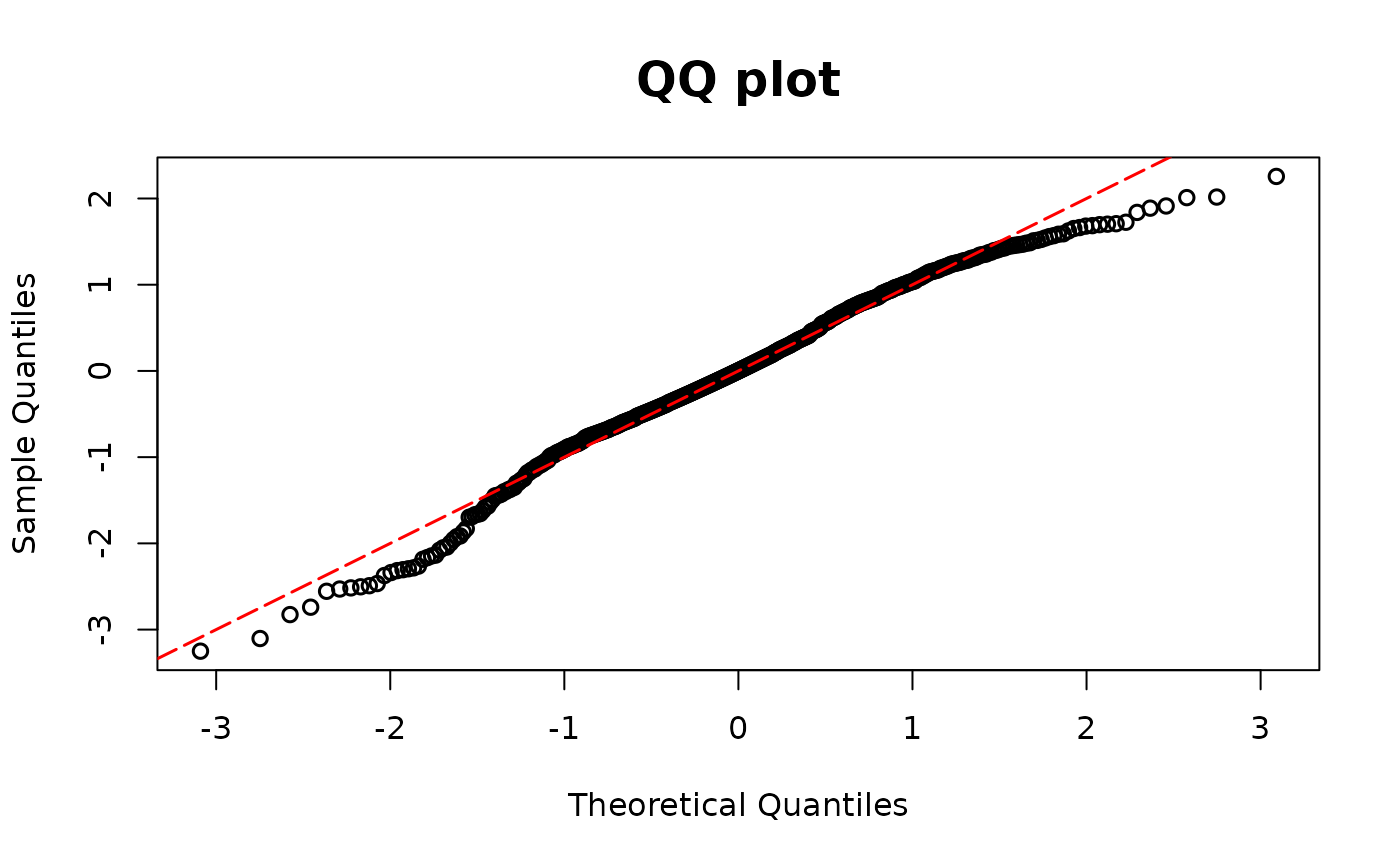
 # Missing covariates
model02 <- glm(y1 ~ x1, family = binomial(link = "logit"))
y2 <- model02$y
fitted2 <- fitted(model02)
resid.bin2 <- dpit_bin(y=y2, prob=fitted2)
# Missing covariates
model02 <- glm(y1 ~ x1, family = binomial(link = "logit"))
y2 <- model02$y
fitted2 <- fitted(model02)
resid.bin2 <- dpit_bin(y=y2, prob=fitted2)