Calculates DPIT residuals with model for semi-continuous outcomes.
dpit_2pm can be used either with model0 and model1 or with part0 and part1 as arguments.
Usage
dpit_2pm(model0, model1, y, part0, part1, plot=TRUE, scale = "normal",
line_args= list(), ...)Arguments
- model0
Model object for 0 outcomes (e.g., logistic regression)
- model1
Model object for the continuous part (gamma regression)
- y
Semicontinuous outcomes.
- part0
Alternative argument to
model0. One can supply the sequence of probabilities \(P(Y_i=0),~i=1,\ldots,n\).- part1
Alternative argument to
model1. One can fit a regression model on the positive data and supply their probability integral transform. Note that the length ofpart1is the number of positive values inyand can be shorter thanpart0.- plot
A logical value indicating whether or not to return QQ-plot
- scale
You can choose the scale of the residuals among
normalanduniform. The default scale isnormal.- line_args
A named list of graphical parameters passed to
graphics::abline()to modify the reference (red) 45° line in the QQ plot. If left empty, a default red dashed line is drawn.- ...
Additional graphical arguments passed to
stats::qqplot()for customizing the QQ plot (e.g.,pch,col,cex,xlab,ylab).
Details
For formulation details on semicontinuous outcomes, see dpit.
In two-part models, the probability of zero can be modeled using a logistic regression, model0,
while the positive observations can be modeled using a gamma regression, model1.
Users can choose to use different models and supply the resulting probabilities of zero and probability integral transforms.
part0 should be the sequence of fitted probabilities of zeros \(\hat{p}_0(\mathbf{X}_i) ,~i=1,\ldots,n\).
part1 should be the probability integral transform of the positive part \(\hat{G}(Y_i|\mathbf{X}_i)\).
Note that the length of part1 is the number of positive values in y and can be shorter than part0.
Examples
library(MASS)
n <- 500
beta10 <- 1
beta11 <- -2
beta12 <- -1
beta13 <- -1
beta14 <- -1
beta15 <- -2
x11 <- rnorm(n)
x12 <- rbinom(n, size = 1, prob = 0.4)
p1 <- 1 / (1 + exp(-(beta10 + x11 * beta11 + x12 * beta12)))
lambda1 <- exp(beta13 + beta14 * x11 + beta15 * x12)
y2 <- rgamma(n, scale = lambda1 / 2, shape = 2)
y <- rep(0, n)
u <- runif(n, 0, 1)
ind1 <- which(u >= p1)
y[ind1] <- y2[ind1]
# models as input
mgamma <- glm(y[ind1] ~ x11[ind1] + x12[ind1], family = Gamma(link = "log"))
m10 <- glm(y == 0 ~ x12 + x11, family = binomial(link = "logit"))
resid.model <- dpit_2pm(model0 = m10, model1 = mgamma, y = y)
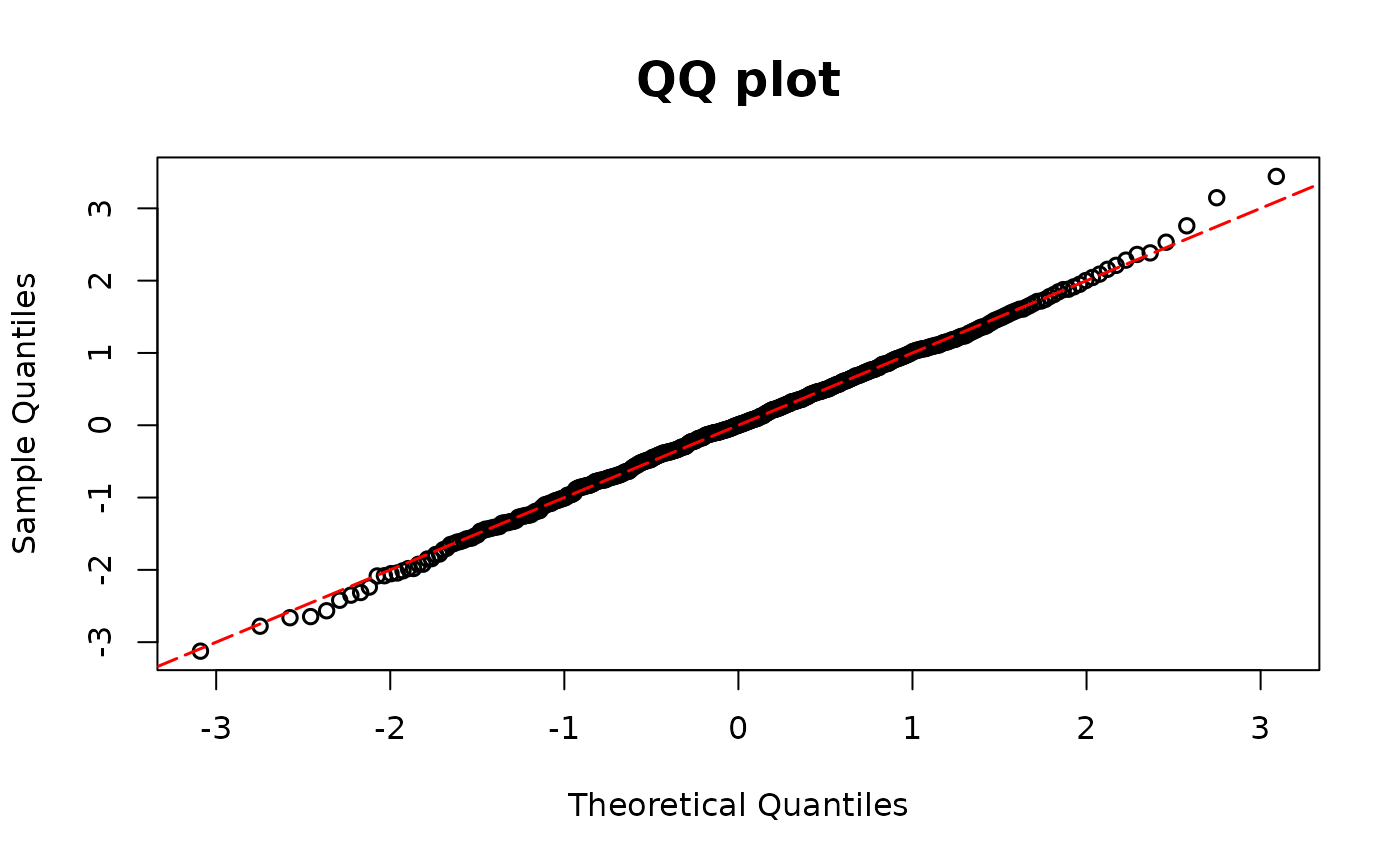 # PIT as input
cdfgamma <- pgamma(y[ind1],
scale = mgamma$fitted.values * gamma.dispersion(mgamma),
shape = 1 / gamma.dispersion(mgamma)
)
p1f <- m10$fitted.values
resid.pit <- dpit_2pm(y = y, part0 = p1f, part1 = cdfgamma)
# PIT as input
cdfgamma <- pgamma(y[ind1],
scale = mgamma$fitted.values * gamma.dispersion(mgamma),
shape = 1 / gamma.dispersion(mgamma)
)
p1f <- m10$fitted.values
resid.pit <- dpit_2pm(y = y, part0 = p1f, part1 = cdfgamma)