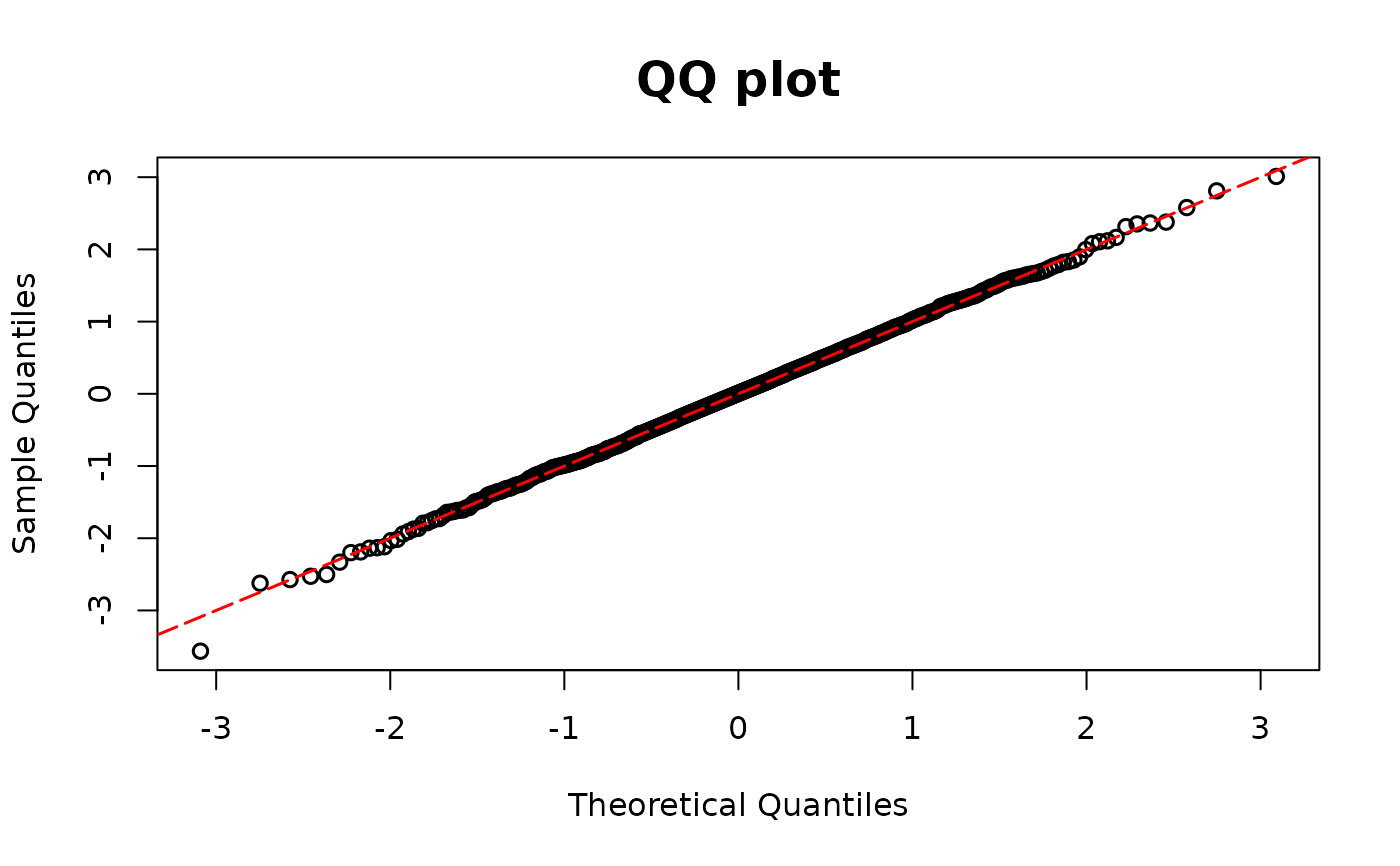
Computes DPIT residuals for regression models with ordinal outcomes
using observed outcomes (y), ordinal outcome levels (level) and their fitted category
probabilities (fitprob).
Usage
dpit_ordi(y, level, fitprob, plot=TRUE, scale="normal", line_args=list(), ...)Arguments
- y
An observed ordinal outcome vector.
- level
The names of the response levels. For instance, c(0,1,2).
- fitprob
A matrix of fitted category probabilities. Each row corresponds to an observation, and column j contains the fitted probability P(Y_i = j).
- plot
A logical value indicating whether or not to return QQ-plot
- scale
You can choose the scale of the residuals among
normalanduniform. The sample quantiles of the residuals are plotted against the theoretical quantiles of a standard normal distribution under the normal scale, and against the theoretical quantiles of a uniform (0,1) distribution under the uniform scale. The default scale isnormal.- line_args
A named list of graphical parameters passed to
graphics::abline()to modify the reference (red) 45° line in the QQ plot. If left empty, a default red dashed line is drawn.- ...
Additional graphical arguments passed to
stats::qqplot()for customizing the QQ plot (e.g.,pch,col,cex,xlab,ylab).
Details
For formulation details on discrete outcomes, see dpit.
Examples
## Ordinal example
library(MASS)
n <- 500
x1 <- rnorm(n, mean = 2)
beta1 <- 3
# True model
p0 <- plogis(1, location = beta1 * x1)
p1 <- plogis(4, location = beta1 * x1) - p0
p2 <- 1 - p0 - p1
genemult <- function(p) {
rmultinom(1, size = 1, prob = c(p[1], p[2], p[3]))
}
test <- apply(cbind(p0, p1, p2), 1, genemult)
y1 <- rep(0, n)
y1[which(test[1, ] == 1)] <- 0
y1[which(test[2, ] == 1)] <- 1
y1[which(test[3, ] == 1)] <- 2
multimodel <- polr(as.factor(y1) ~ x1, method = "logistic")
y1 <- multimodel$model[,1]
lev1 <- multimodel$lev
fitprob1 <- fitted(multimodel)
resid.ord <- dpit_ordi(y=y1, level=lev1, fitprob=fitprob1)